I attended the Evolution 2014 meeting a few months ago in Raleigh, NC, and presented a poster on Phenoscape’s curation effort: “Moving the mountain: How to transform comparative anatomy into computable anatomy?”, with coauthors A. Dececchi, N. Ibrahim, H. Lapp, and P. Mabee. In this work, we assessed the efficiency of our workflow for the curation of evolutionary phenotypes from the matrix-based phylogenetic literature. We identified the bottlenecks and areas of improvement in data preparation, phenotype annotation, and ontology development. Gains in efficiency, such as through improved community data practices and development of text-mining tools, are critical if we are to translate evolutionary phenotypes from an ever-growing literature. The poster was well received and several researchers at the meeting were interested in learning more about open source tools for phenotype annotation.
Phenoscape makes a splash at SVP, TDWG
January 10, 2014In an effort to expand the user community and to demonstrate what is possible using our infrastructure, members of the Phenoscape team gave multiple presentations across two continents on our recent developments. In late October Paula Mabee gave an invited presentation on mapping phenotypes across phylogenies at the Muséum national d’Histoire naturelle in Paris. This was followed by presentations at the 73rd annual meeting of the Society of Vertebrate Paleontology (SVP) in Los Angeles and the 2013 meeting of the Taxonomic Database Working Group (TDWG) in Florence, Italy. Phenoscape had a significant presence at SVP with both a poster presented by Alex Dececchi demonstrating our progress in generating supermatrices from our annotations as well as a talk given by collaborator Karen Sears, using EQ supermatrices from Phenoscape fin/limb data to examine integration patterns across the fin to limb transition. Karen’s talk marks the first of the collaborations coming out of our 2013 San Francisco workshop. It also showed how data from Phenoscape can drive independent projects and is easily integrated with existing phylogenetic and statistical tool such as Mesquite and various R modules. The talks and poster were well received, with numerous researchers inquiring on how they could incorporate Phenoscape or use ontology based annotations.
Society of Vertebrate Paleontology Annual Meeting 2012 (Raleigh, NC)
November 27, 2012The Phenoscape project had a strong presence at the largest Vertebrate Paleontology/Comparative Anatomy conference in the world this year, the Society of Vertebrate Paleontology annual meeting. In one of the large conference halls, and in front of a packed audience, I gave a talk on the history, goals and background of the Phenoscape project (“Phenoscape: A New Anatomical Ontology of Vertebrates”). The authorship also included Paul Sereno, Paula Mabee, Todd Vision and Hilmar Lapp. The talk was well received, and several attendees expressed great interest in our work. The difficult part now is to make sure this first spark of interest is maintained – this can be difficult when the community has not been exposed to ontologies before and the project appears to be so different from anything they have done before – but we’ll do our best to stay in contact with those people that expressed strong interest.
Alex Dececchi presented a poster on Phenoscape at the same conference (Phenoscape: bridging the gap between fossils and genes – his co-authors were J. Balhoff, W. Dahdul, N. Ibrahim, H. Lapp, P. Midford, P. Sereno, T. Vision, M. Westerfield, P Mabee and D. Blackburn), making sure that even those that could not attend the talk would get an opportunity to learn more about our exciting work.
Nizar Ibrahim, University of Chicago
DILS 2012
August 28, 2012In June I had the opportunity to attend DILS 2012 (Data Integration in the Life Sciences), at the University of Maryland in College Park. I presented a poster on Phenoscape, “The Phenoscape Knowledgebase: Integrating phenotypic data across taxonomy, from biodiversity to developmental genetics”. The poster highlighted some of the new directions the Phenoscape project is heading, such as broadening taxonomic coverage and adoption of semantic web technologies. DILS was a small conference but had several talks discussing the applications of ontologies to biological data. I’m looking forward to DILS 2013 in Montreal, in conjunction with ICBO and the Canadian Semantic Web conference.
Notes from TDWG 2011
November 22, 2011Last month, I (Jim Balhoff) and Hilmar Lapp attended the Biodiversity Information Standards meeting (TDWG 2011), in New Orleans. As a representative of both Phenoscape and the Hymenoptera Anatomy Ontology project, I presented a poster, with co-authors Matt Yoder and Andy Deans, detailing an OWL model showing the explicit semantics of linking an Entity–Quality (EQ) phenotype to evolutionary character matrix data and taxonomic specimens. While EQ can be thought of as simple ontological tags on descriptive data, modeling phenotypes within a more explicit logical framework allows us to make use of more powerful automated reasoning. It also provides a consistent interpretation for EQs across projects annotating phenotypes (for example, Phenoscape and HAO). 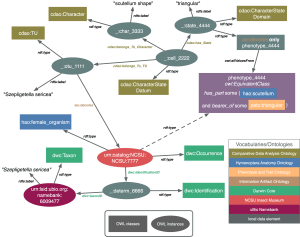
Of particular relevance to our poster was another presented by Cam Webb. Cam has created an OWL-compatible version of Darwin Core which can be used to describe specimen metadata in RDF. We made similar use of Darwin Core in our poster, but we are looking into adopting Cam’s Darwin-SW for this part of the model.
Overall there was a lot of interest in semantic technologies at TDWG, ranging from the initial meeting of an RDF/OWL working group to other projects that are not using semantic technologies but seem well suited for RDF.
Notes from ISWC 2011
November 3, 2011Last week, I attended the 10th International Semantic Web Conference (ISWC) in Bonn, Germany. A tremendous variety of sophisticated work is going on both in academia and industry to improve the technology for, and take advantage of, the ever-growing network of data and concepts published, through open standards, on the web.
You might say it is the best of times and the worst of times for semantic web enthusiasts, in that reasoning and query engines that can be used on large collections of RDF have in the last few years become a reality (one of the Challenge Tracks provided contestants with a *billion* triples to work with). But some see clouds on the horizon. The web search titans (Bing, Google and Yahoo!) are now pushing schema.org, a microformat and vocabulary standard for web content that some worry may threaten the development of richer semantic web technology. Still, most treated the news positively, happy to know that these organizations now seem to agree on the importance of semantics. In fact, Yahoo! described at the conference how they are trying to build a “Web of Objects” that takes advantage of scheme.org, together with more extensive internal vocabularies, to regroup knowledge pieces that are scattered around the Web.
Conference chair Natasha Noy showed a revealing pair of tag clouds comparing the abstracts from the first year of the conference in 2001 to today — the terms “semantic” and “web” have shrunk in importance and “data” is now king! 
Ivan Herman’s blog gives a good sampling of the flavor of talks presented at the meeting. I especially enjoyed the Industry Track, since these applications are less familiar to me than the academic/scientific ones, and I was particularly impressed by the importance of semantic technologies to the news media and other content industries. These technologies are being deployed by news organizations with great enthusiam (e.g. the BBC). I also came away with a strong sense that semantic technologies are helping to create demand, and drive a revolution in the use of, Open Government Data; there were a number of demonstrations of useful real-world applications, particularly to environmental monitoring.
With my Phenoscape hat on, I attended a Linked Open Data for Science (LISC) satellite workshop prior to the main conference. The event included both presentations and discussions from a variety of perspectives about the opportunities and challenges of this new technology. A diversity of fields were represented (social science, linguistics, geosciences, biomedicine, etc.). But, it is clear that uptake of linked open data as an alternative means of publication is still in its infancy within the sciences. This despite the fact that the bioinformatics data centers account for nearly a quarter of the real estate in the famous linked data cloud diagram. Some of the most exciting opportunities, in my opinion, come from the ability to allow radically decentralized data publication, and this is something that we might wish to pilot in a modestly distributed data curation environment like Phenoscape. Another observation: I was surprised to discover at the meeting how much the utility of the linked data cloud (and, by extension, the semantic web) depend on the social convention by which everyone provides links into a relatively small number of large ‘concept repositories’ like DBPedia (which was originally a Master’s project, BTW).
The breakout discussion sessions at LISC highlighted how scientific practice will place difficult demands on linked data with respect to provenance, context, granularity, distributed authority, etc. This resonated with the message of our own contribution to the workshop, which outlined some of the particular challenges in making context-dependent links between scientific objects, when the descriptions of those objects are scattered across different resources, and when the similarities between objects are spread weakly over many properties [1]. Another important question that hit home for a number of us coming from the bioinformatics and biodiversity informatics world is how scientists are going to be able to take advantage of the innovations now going on in the commercial sector (including some of the exhibitors at the main conference) within the constraints and DIY culture of small individual university-based research grants.
There is no denying the explosion in linked data resources out there (comparisons of the growth in the cloud diagram are about as common as graphs showing the growth in sequence data at a biology conference). But another recurrent theme of the meeting was that unfortunately much of that content is missing semantics (i.e. a lack of use or availability of ontologies for many concepts, and lack of links between content at different endpoints), and generating semantically annotated triples needs to be easier that it currently is (a message certainly relevant to those of us developing curation tools).
One of the keynotes, from Frank van Harmelen, generated quite a bit of buzz. He looked back on 10 years of the semantic web, asking what theoretical principles we can learn from the experience so far, and his annotated slides are well worth a look.
The conference was a great mix of different formats. In addition to the keynotes and regular talks, there are a host of workshops and tutorials, challenges, panel discussions (including one billed as a ‘Death Match’), and even a special competition for the best “Outrageous Ideas”. The winner of that one was a proposal to bring linked data to the non-networked portion of humanity. A particularly nice feature of the meeting was the ‘Minute Madness’ preceding the poster session in which each of the poster presenters gave a short timed pitch with to all the attendees – it was a very entertaining and informative way to ‘see’ every poster and allowed everyone to quickly pick out which ones to hit during the session.
For more, see the excellent day-by-day summary of the meeting from Juan Sequeda, where there are links to all the winning presentations and challenge entries. [Ironically, the conference website is down temporarily while it is being moved, so come back later if the links to the papers hang]. The next ISWC will be November 11-15, 2012 in Boston.
Reference:
[1] Vision T, Blake J, Lapp H, Mabee P, Westerfield M (2011) Similarity Between Semantic Description Sets: Addressing Needs Beyond Data Integration, in Proceedings of the First International Workshop on Linked Science, Bonn, Germany, October 24, 2011, Tomi Kauppinen, Line C. Pouchard, Carsten Kessler (eds), published in CEUR Workshop Proceedings, Volume 783.
ICBO 2011
August 11, 2011Jim Balhoff and I recently attended the International Conference on Biomedical Ontology (ICBO) held 26-30 July in Buffalo, NY. The conference focused on the use and development of ontologies in the biological and biomedical domains. Of particular interest to Phenoscape were the workshops and tutorials held during the two days before the main conference. Topics included ontology integration, Common Logic, ontology development tools, and using OBO and OWL formats for ontology development and reasoning.
We presented talks at the Facilitating Anatomy Ontology Interoperability workshop. Jim’s talk was on representing taxa as individuals in OWL, an alternative to the common representation of taxa as classes, which facilitates annotation of phenotypic data involving polymorphism and evolutionary reversals. I presented a lightning talk on the anatomy ontology synchronization requirements for linking evolutionary and model organism phenotypes. Other presentations from the workshop are available here. We also presented a poster describing the reasoning used in the Phenoscape Knowledgebase.
The main conference included interesting talks on a broad range of topics including the application of ontologies to proteins, diseases, biological mechanisms, and electronic health records. Presentations can be downloaded here.
CSHALS 2011
March 9, 2011I recently attended the Conference on Semantics in Healthcare and Life Sciences (CSHALS), in Cambridge, MA. The CSHALS meeting was a change for me in that it’s much more healthcare-oriented than other venues in which I’ve presented work from Phenoscape. This was a great opportunity to see how far the healthcare community has pushed semantic web technologies, and also to become more familiar with some of the more commercial packages which are available for storing and querying very large knowledgebases based on RDF (for example, AllegroGraph and Gruff from Franz, Inc., and Sentient Knowledge Explorer from IO Informatics). A particularly interesting talk was the keynote by Toby Segaran, of Metaweb Technologies, advocating semantic techniques as a more agile approach to data. Slideshows from the conference presentations are available for download here, including my own.



 Posted by wdahdul
Posted by wdahdul 